TASSEL软件下载
- 2026-05-13 13:01:48
TASSEL软件下载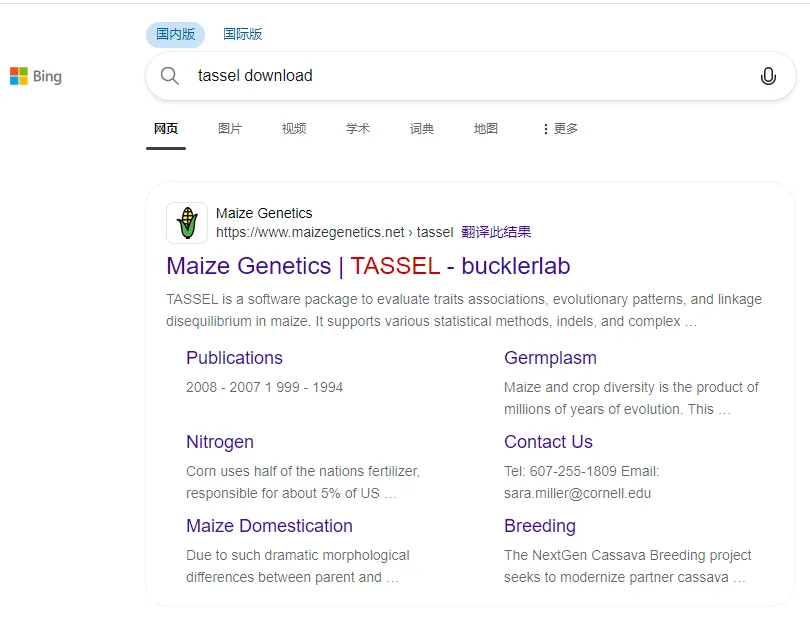
大家好,我是邓飞,这一篇主要是灌水,因为有小伙伴问我Tassel的安装包,因为tassel下载网络不太好:

点击之后:

用我的方法吧!
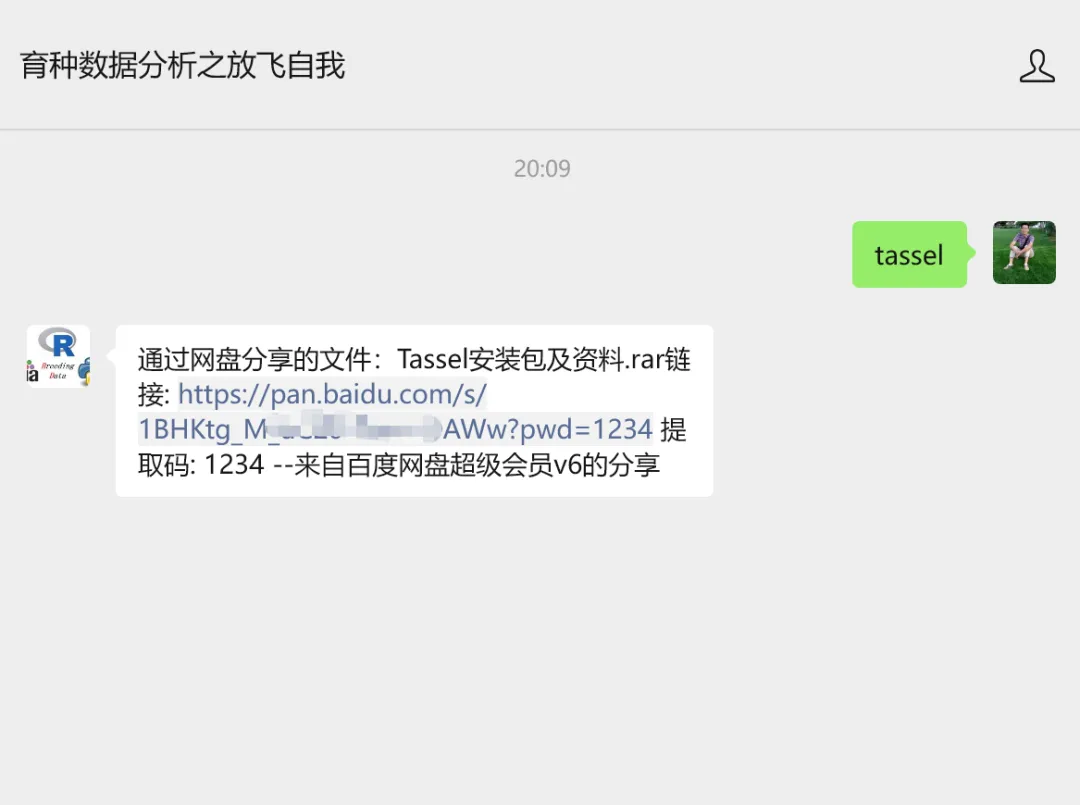
TASSEL配套数据和代码,公众号回复:tassel,获得下载链接:

相关教程:
第二篇:对基因型数据质控:缺失质控,maf质控,hwe质控,样本质控
第六篇:就是这篇。
1. TASSEL的GLM和MLM分析结果
质控后的plink数据和表型数据:
 「GLM的GWAS分析结果:」
「GLM的GWAS分析结果:」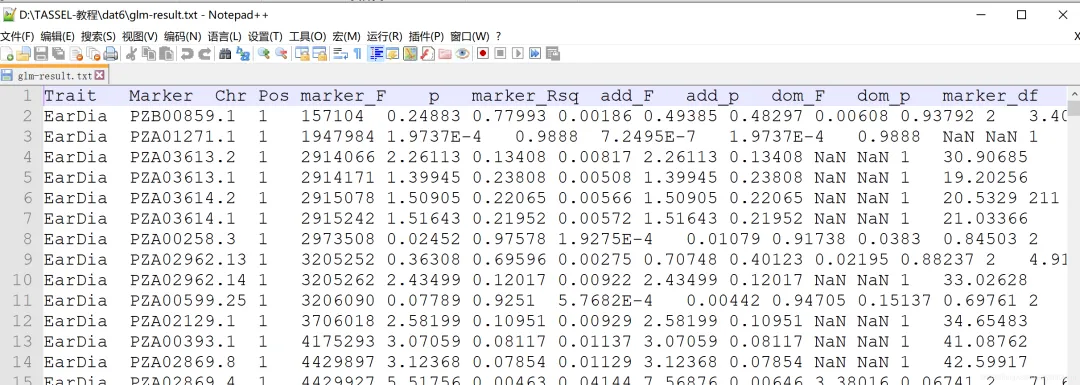 「MLM的GWAS分析结果:」
「MLM的GWAS分析结果:」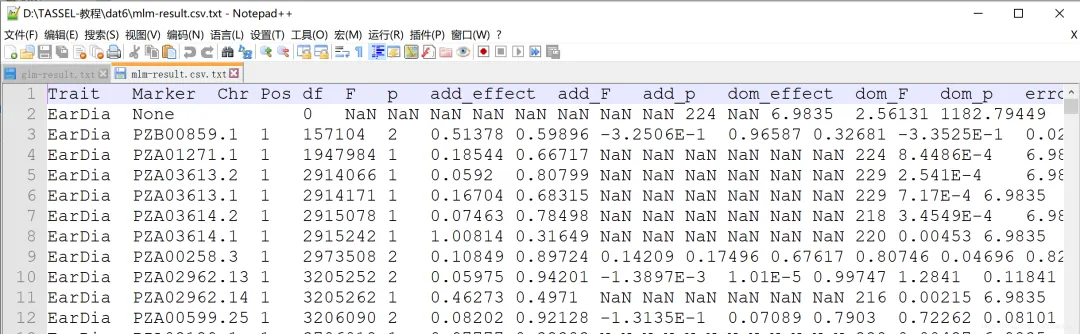
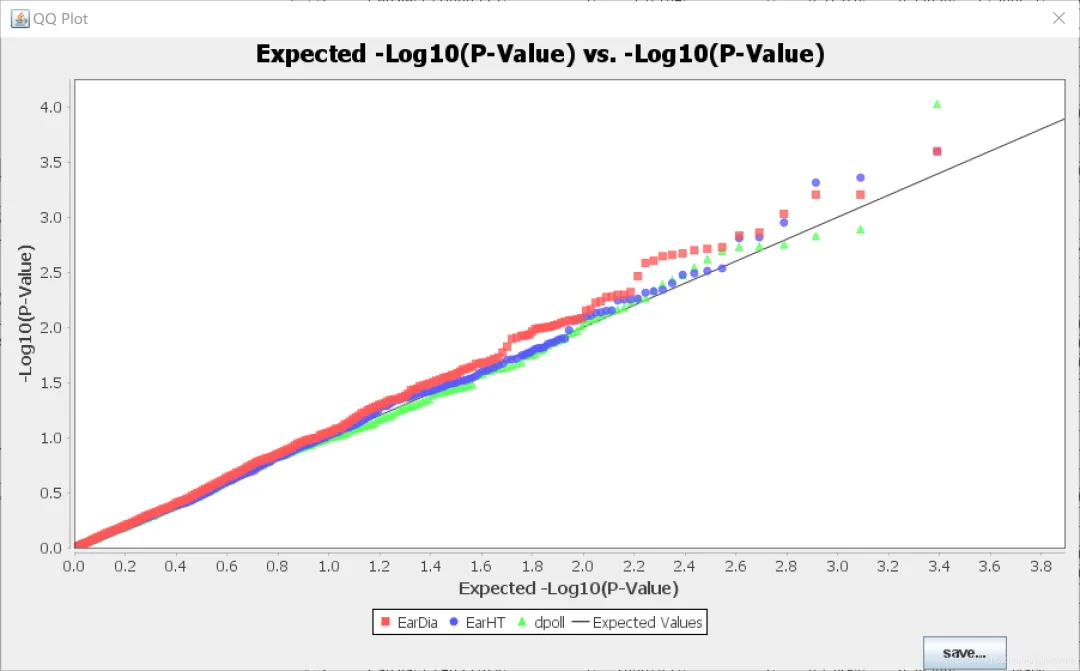
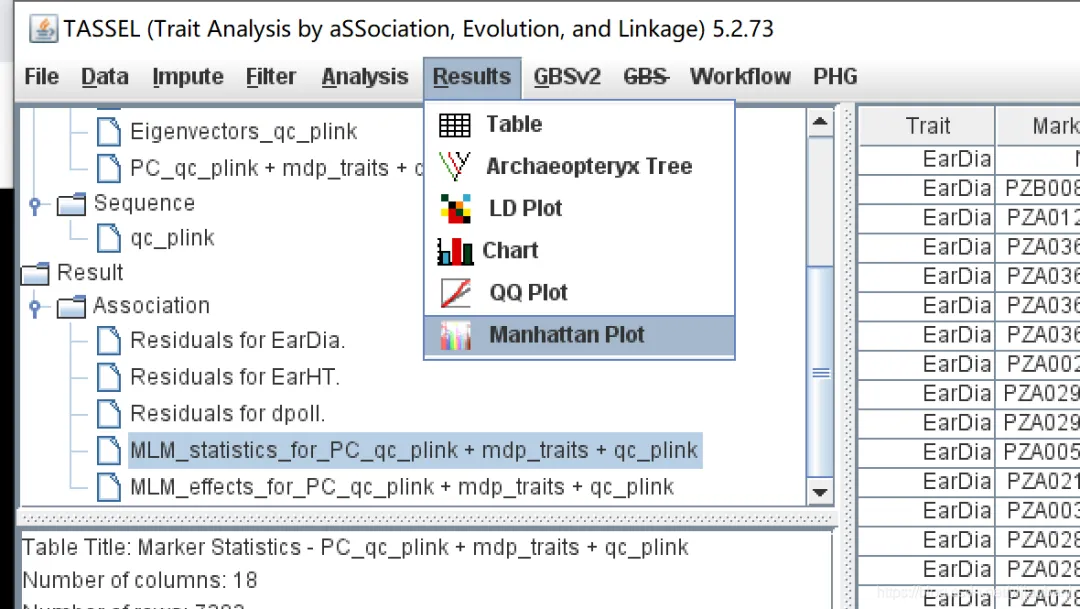
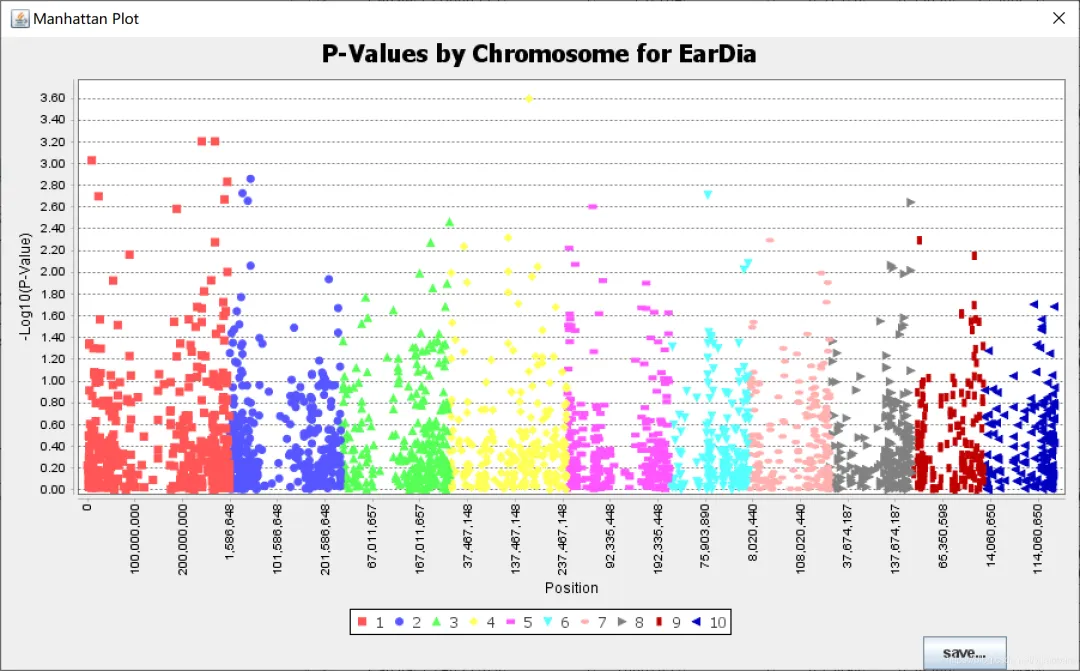
2. TASSEL中的可视化
TASSEL有对结果进行可视化的模块,包括qq图和曼哈顿图,但是图不方便调整。这里用TASSEL的分析结果,使用R语言进行绘制qq图和曼哈顿图。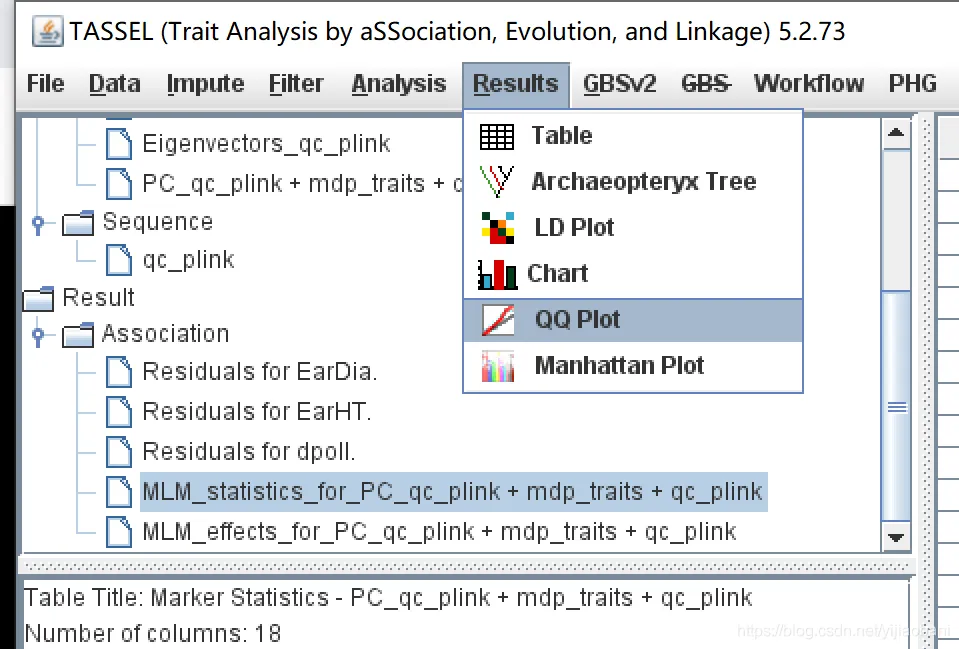



3. R语言包安装及载入
需要用到:
qqmantidyversedata.table
下面代码,会判断是否有这三个包,如果没有,就自动安装。然后载入软件包。
if(!require(data.table)) install.packages("data.table")if(!require(qqman)) install.packages("qqman")if(!require(tidyverse)) install.packages("tidyverse")library(qqman)library(tidyverse)library(data.table)4. GLM模型GWAS结果可视化
results_log = fread("glm-result.txt")dim(results_log)head(results_log)select = dplyr::selecttable(results_log$Trait)结果:
> table(results_log$Trait) dpoll EarDia EarHT 2460 2460 2460数据中共有三个性状,可以选择一个性状,进行可视化。
d1 = results_log %>% filter(Trait == "dpoll") %>% select(Chr,Marker,Pos,p)head(d1)summary(d1)d1 = d1 %>% drop_na(p)summary(d1)注意,有些P值是NA,在作图时会报错,这里将其移除。
整理后的结果:
> summary(d1) Chr Marker Pos p Min. : 1.0 Length:2460 Min. : 139753 Min. :0.0000 1st Qu.: 2.0 Class :character 1st Qu.: 43868061 1st Qu.:0.1236 Median : 4.0 Mode :character Median :128423374 Median :0.3911 Mean : 4.7 Mean :120382976 Mean :0.4165 3rd Qu.: 7.0 3rd Qu.:175628840 3rd Qu.:0.6743 Max. :10.0 Max. :298413352 Max. :0.9996作图代码:
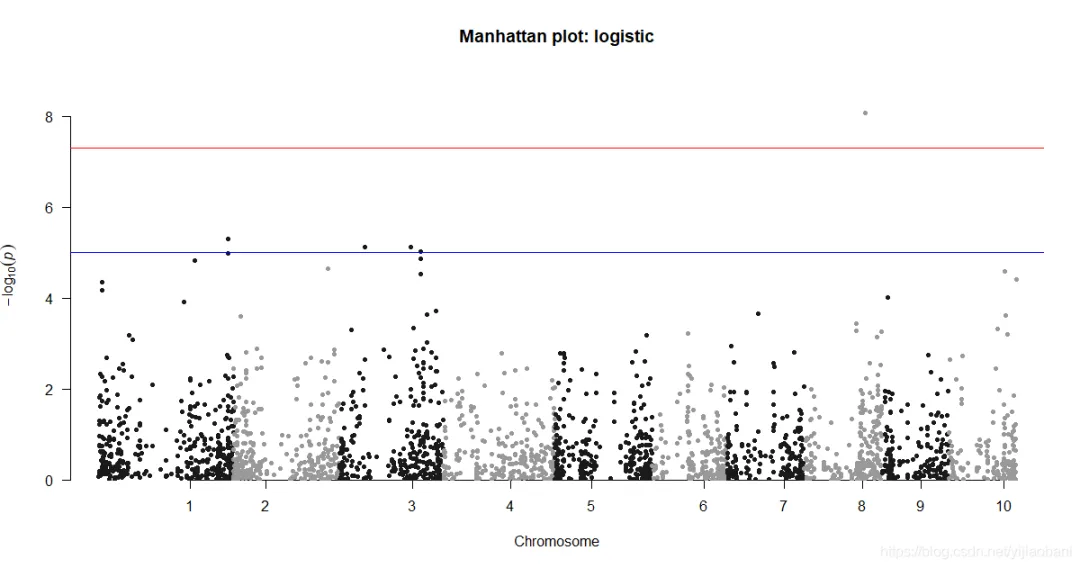
manhattan(d1,chr="Chr",bp="Pos",p="p",snp="Marker", main = "Manhattan plot: logistic")tiff("y1-曼哈顿图.tiff")manhattan(d1,chr="Chr",bp="Pos",p="p",snp="Marker", main = "Manhattan plot: logistic")dev.off()qq(d1$p, main = "Q-Q plot of GWAS p-values : log")tiff("y1-QQ图.tiff")qq(d1$p, main = "Q-Q plot of GWAS p-values : log")dev.off()「曼哈顿图:」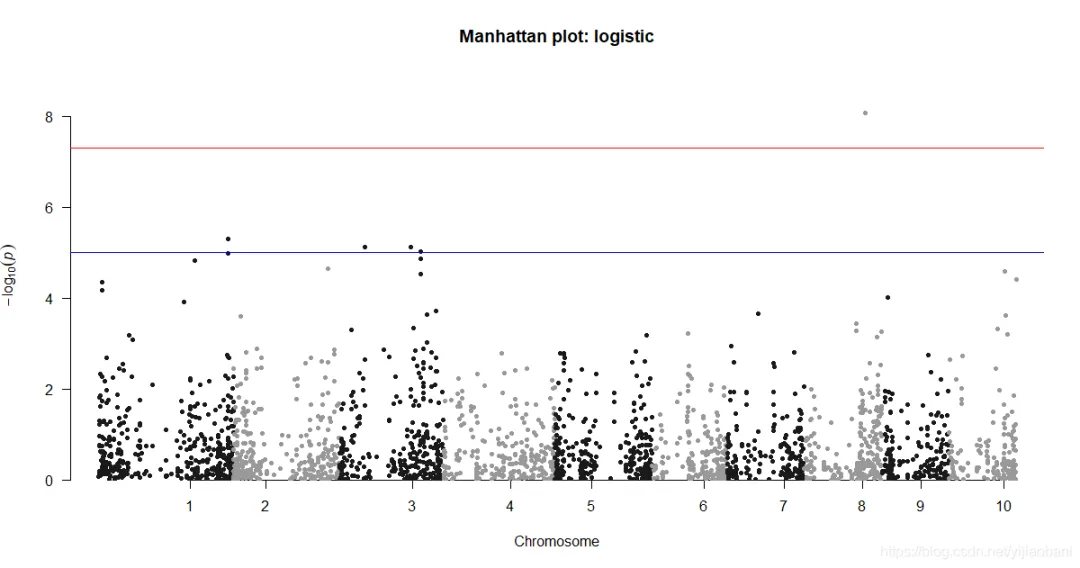
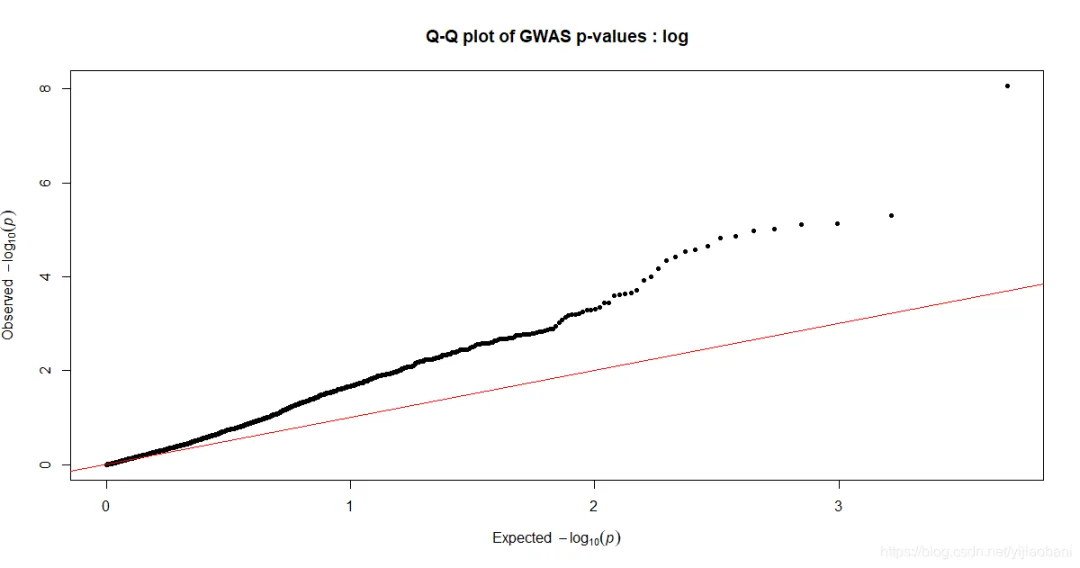
「QQ图:」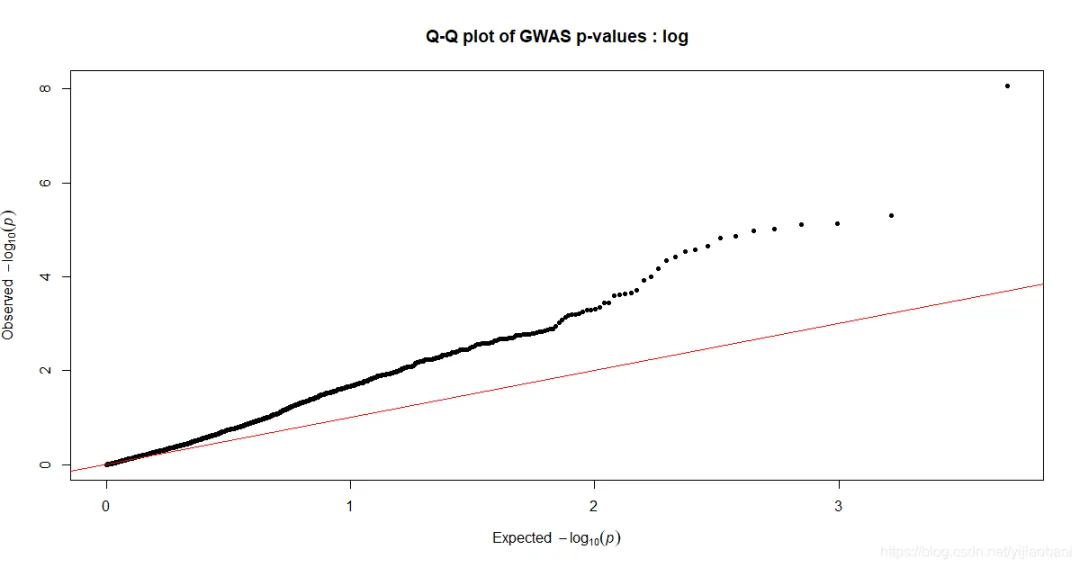 其它两个性状的作图代码:
其它两个性状的作图代码:
d2 = results_log %>% filter(Trait == "EarDia") %>% select(Chr,Marker,Pos,p)head(d2)summary(d2)d2 = d2 %>% drop_na(p)summary(d2)tiff("y2-曼哈顿图.tiff")manhattan(d2,chr="Chr",bp="Pos",p="p",snp="Marker", main = "Manhattan plot: logistic")dev.off()tiff("y2-QQ图.tiff")qq(d2$p, main = "Q-Q plot of GWAS p-values : log")dev.off()d3 = results_log %>% filter(Trait == "EarHT") %>% select(Chr,Marker,Pos,p)head(d3)summary(d3)d3 = d3 %>% drop_na(p)summary(d3)tiff("y3-曼哈顿图.tiff")manhattan(d3,chr="Chr",bp="Pos",p="p",snp="Marker", main = "Manhattan plot: logistic")dev.off()tiff("y3-QQ图.tiff")qq(d3$p, main = "Q-Q plot of GWAS p-values : log")dev.off()将整理后的不同性状的结果保存到本地:
fwrite(d1,"y1_result.csv")fwrite(d2,"y2_result.csv")fwrite(d3,"y3_result.csv")5. MLM模型GWAS结果可视化
读取数据,提取性状,去掉P值为缺失的行:
library(qqman)library(data.table)results_log = fread("mlm-result.txt", head=TRUE)dim(results_log)head(results_log)library(tidyverse)select = dplyr::selecttable(results_log$Trait)d1 = results_log %>% filter(Trait == "dpoll") %>% select(Chr,Marker,Pos,p)head(d1)summary(d1)d1 = d1 %>% drop_na(p)summary(d1)「曼哈顿图:」

「QQ图:」

其它两个作图代码:
d2 = results_log %>% filter(Trait == "EarDia") %>% select(Chr,Marker,Pos,p)head(d2)summary(d2)d2 = d2 %>% drop_na(p)summary(d2)tiff("y2-曼哈顿图.tiff")manhattan(d2,chr="Chr",bp="Pos",p="p",snp="Marker", main = "Manhattan plot: logistic")dev.off()tiff("y2-QQ图.tiff")qq(d2$p, main = "Q-Q plot of GWAS p-values : log")dev.off()d3 = results_log %>% filter(Trait == "EarHT") %>% select(Chr,Marker,Pos,p)head(d3)summary(d3)d3 = d3 %>% drop_na(p)summary(d3)tiff("y3-曼哈顿图.tiff")manhattan(d3,chr="Chr",bp="Pos",p="p",snp="Marker", main = "Manhattan plot: logistic")dev.off()tiff("y3-QQ图.tiff")qq(d3$p, main = "Q-Q plot of GWAS p-values : log")dev.off()6. 完整代码汇总
「GLM的可视化代码:」
## 对TASSEL GLM 模型可视化if(!require(data.table)) install.packages("data.table")if(!require(qqman)) install.packages("qqman")if(!require(tidyverse)) install.packages("tidyverse")library(qqman)library(tidyverse)library(data.table)results_log = fread("glm-result.txt")dim(results_log)head(results_log)select = dplyr::selecttable(results_log$Trait)d1 = results_log %>% filter(Trait == "dpoll") %>% select(Chr,Marker,Pos,p)head(d1)summary(d1)d1 = d1 %>% drop_na(p)summary(d1)manhattan(d1,chr="Chr",bp="Pos",p="p",snp="Marker", main = "Manhattan plot: logistic")tiff("y1-曼哈顿图.tiff")manhattan(d1,chr="Chr",bp="Pos",p="p",snp="Marker", main = "Manhattan plot: logistic")dev.off()qq(d1$p, main = "Q-Q plot of GWAS p-values : log")tiff("y1-QQ图.tiff")qq(d1$p, main = "Q-Q plot of GWAS p-values : log")dev.off()d2 = results_log %>% filter(Trait == "EarDia") %>% select(Chr,Marker,Pos,p)head(d2)summary(d2)d2 = d2 %>% drop_na(p)summary(d2)tiff("y2-曼哈顿图.tiff")manhattan(d2,chr="Chr",bp="Pos",p="p",snp="Marker", main = "Manhattan plot: logistic")dev.off()tiff("y2-QQ图.tiff")qq(d2$p, main = "Q-Q plot of GWAS p-values : log")dev.off()d3 = results_log %>% filter(Trait == "EarHT") %>% select(Chr,Marker,Pos,p)head(d3)summary(d3)d3 = d3 %>% drop_na(p)summary(d3)tiff("y3-曼哈顿图.tiff")manhattan(d3,chr="Chr",bp="Pos",p="p",snp="Marker", main = "Manhattan plot: logistic")dev.off()tiff("y3-QQ图.tiff")qq(d3$p, main = "Q-Q plot of GWAS p-values : log")dev.off()fwrite(d1,"y1_result.csv")fwrite(d2,"y2_result.csv")fwrite(d3,"y3_result.csv")「MLM的可视化代码:」
## 对TASSEL GLM 模型可视化library(qqman)library(data.table)results_log = fread("mlm-result.txt", head=TRUE)dim(results_log)head(results_log)library(tidyverse)select = dplyr::selecttable(results_log$Trait)d1 = results_log %>% filter(Trait == "dpoll") %>% select(Chr,Marker,Pos,p)head(d1)summary(d1)d1 = d1 %>% drop_na(p)summary(d1)manhattan(d1,chr="Chr",bp="Pos",p="p",snp="Marker", main = "Manhattan plot: logistic")tiff("y1-曼哈顿图.tiff")manhattan(d1,chr="Chr",bp="Pos",p="p",snp="Marker", main = "Manhattan plot: logistic")dev.off()qq(d1$p, main = "Q-Q plot of GWAS p-values : log")tiff("y1-QQ图.tiff")qq(d1$p, main = "Q-Q plot of GWAS p-values : log")dev.off()d2 = results_log %>% filter(Trait == "EarDia") %>% select(Chr,Marker,Pos,p)head(d2)summary(d2)d2 = d2 %>% drop_na(p)summary(d2)tiff("y2-曼哈顿图.tiff")manhattan(d2,chr="Chr",bp="Pos",p="p",snp="Marker", main = "Manhattan plot: logistic")dev.off()tiff("y2-QQ图.tiff")qq(d2$p, main = "Q-Q plot of GWAS p-values : log")dev.off()d3 = results_log %>% filter(Trait == "EarHT") %>% select(Chr,Marker,Pos,p)head(d3)summary(d3)d3 = d3 %>% drop_na(p)summary(d3)tiff("y3-曼哈顿图.tiff")manhattan(d3,chr="Chr",bp="Pos",p="p",snp="Marker", main = "Manhattan plot: logistic")dev.off()tiff("y3-QQ图.tiff")qq(d3$p, main = "Q-Q plot of GWAS p-values : log")dev.off()fwrite(d1,"y1_result.csv")fwrite(d2,"y2_result.csv")fwrite(d3,"y3_result.csv")另外,之前推荐的平台,发现也上线了GWAS模块,但是有点单一,简单是很简单:

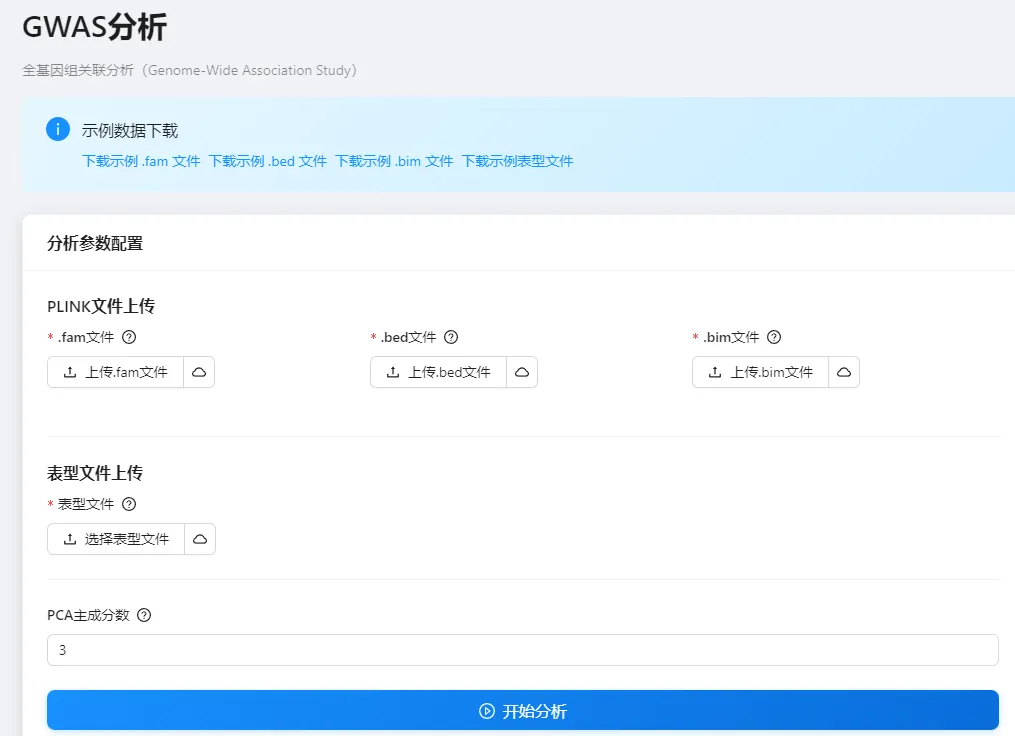
1,上传plink二进制文件
2,上传表型数据
3,指定pca个数
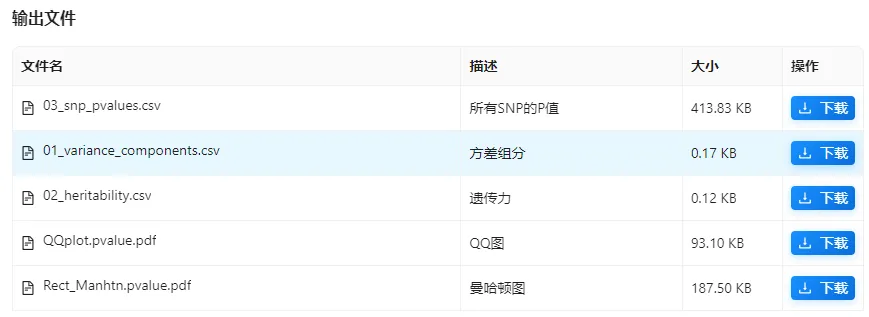
4,查看结果,包括GWAS的Pvalue结果(包括pve解释百分比、QQ图、曼哈顿图)

平台地址:https://asreml.cn/gwas-analysis
本文来自网友投稿或网络内容,如有侵犯您的权益请联系我们删除,联系邮箱:wyl860211@qq.com 。
